




We will evaluate our AdaLab framework on a biomedically important application area: modelling the yeast diauxic shift. When yeast (Saccharomyces cerevisiae) is grown on glucose with oxygen it first produces ethanol, and when the glucose is exhausted it reorganises (shifts) its metabolism to grow using the ethanol it previously produced (Dickinson, 1999). This reorganisation of metabolism is known as the diauxic shift.
In biology it is used as a 'model' for general cellular reorganisation, and has significant medical applications in the understanding of ageing and cancer. In cancer the Warburg effect (a metabolic shift to lactic acid from glucose - lactic acid is the equivalent in animals of ethanol) is typically observed in solid tumours, and is believed to partly explain tumour growth (Gatenby and Gawlinski, 2003). In ageing a metabolic shift away from lactic acid is related to the cell’s stress response, and this is associated with increased cell life-span (Kenyon, 2010).
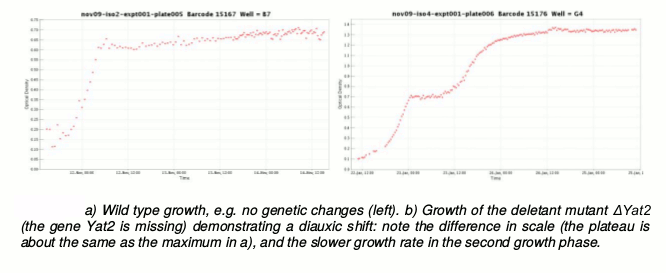
We have established a robust laboratory automation protocol for a high-throughput assay for growth curves. This enables Eve to observe qualitative and quantitative differences in the batch growth in yeast strains. In particular we have good statistical methods that recognise biphasic growth, i.e. diauxic shifts in culture (Fig. 5). These have enabled us to discover novel mutants with aberrant diauxic shifts.
Eve can grow specific yeast strains under specific environmental conditions and make a quantitative measurement of any diauxic shift. The yeast strains will typically have one or more specific genes deleted. The main environmental variations will consist of a defined set of compounds (e.g. glucose, ammonia, etc.) and concentrations (e.g. 1 molar). It is also possible to change the temperature and oxygen content. In addition it is possible to add approved drugs (e.g. aspirin) that are known to interact with specific enzymes. Such high-throughput protocols will be used as the main experimental test of the hypotheses generated by ML and collaborative human scientists. The objective measure of success will be the accuracy of the developed models. These accuracies will be objectively verified by experimental observation of model predictions.
The AdaLab framework requires the OUK knowledge model in order to efficiently record and reason with uncertain knowledge. OUK will enable the recording of verified and unverified knowledge in a consistent manner through the utilisation of probability theory. OUK will import relevant classes from existing biomedical ontologies (i.e. Gene Ontology, Protein Ontology, etc.) and employ the HELO framework to represent biomedical knowledge about the diauxic shift.
We will develop OUK to support the key modules of the AdaLab framework:
Probabilistic reasoning module. OUK will import HELO classes for the representation of data and knowledge items and associated probabilities, and add rules for probabilistic reasoning, i.e. for assigning prior probabilities.
Scientific models and domain knowledge about yeast relevant to cancer and ageing. OUK terms will form a language to declaratively represent research hypotheses that are testable by Eve, and also suitable for ML algorithms, e.g. genes, regulation, conditions, prior distribution over the state of yeast cells. Hypotheses will be represented in a common semantic form as triplets predicate(entity1, entity2), where predicate is the defined in OUK relation between the entities (these also have to be defined in OUK or other bio-ontologies). Complex hypotheses will be formed by the combination of simple statements using logical operators.
ML module. This will import relevant classes from OntoDM (an Ontology of Data Mining, the Coordinator of the proposal is one of the developers) (Panov et. al, 2009), DMOP (Data Mining Ontology) and Expose (Vanschoren and Soldatova, 2010) to provide a descriptive language and to support the recording of the ML run by Eve.
Communication mechanism between human and robot scientists module. We will develop protocols for communications between robot and human scientists based on WSMO (Web Services Modelling Ontology) (Roman et al, 2005), and use of terms defined in OUK. OUK will import the relevant classes from WSMO, and define the modes of communications between humans and robots, message formats, and typical contents of messages. OUK will serve as foundational domain knowledge model for the communication mechanism by defining terms for the communication between robot and human scientists, e.g. task, experimental condition, to change X, to cancel Y; and modes of such communications, e.g. a text message to a mobile phone, email, images from a web camera. OUK will specify Eve’s behaviour: states and transitions. Currently Eve outputs experimental results in a form suitable for understanding by humans, but initial and intermediate states are not recorded in a human-friendly form. We will provide a simple communication mechanism based on the widely accepted WSMO framework. This was developed for communication between web services, but it also facilitates inter-operation between various communities and user groups.

We will extend the capabilities of Robot Scientists, and to enable their better integration with human scientists – the AdaLab framework. At a high level the AdaLab framework will keep track of the current state of the research using the OUK logical framework, suggest what research goals are most interesting, formulate hypotheses towards the most important goals, design experiments to test these hypotheses, and perform suitable statistical analysis. The AdaLab framework will unite the super-human abilities of computers/robots (high inference speed, perfect recall, accuracy of movement) with the creativity and intellectual flexibility of human scientists. To achieve this we will:
Integrate Eve with the developed knowledge representation. Currently our Robot Scientists Eve uses ad hoc relational databases and files to access bioinformatics facts - taken from external bioinformatics databases and the scientific literature. This situation is unsatisfactory and not scalable. We will develop a general scalable solution based on ontologies and use of LOD (Linked Open Data) repositories. This will enable large amounts of heterogeneous scientific data to be integrated and used for reasoning by Robot Scientists.
Integrate Eve with the developed ML and inference methods. Adam used special purpose bioinformatics software and propositional ML to generate hypotheses. The extension to using SRL will enable richer and more general hypotheses to be generated. The newly developed SRL hypothesis generation methods and active learning will enable Eve to more economically form and test hypotheses using principled methods.
Evaluate AdaLab by application to the S. cerevisiae diauxic shift. To evaluate the utility of the proposed AdaLab framework we will demonstrate it working using Eve and human experts in yeast biology, with application to the yeast diauxic shift. We predict that by the end of the project AdaLab will be able to outperform human scientists by more than 20% in this limited but important scientific area, providing an objective evaluation of success. Furthermore, publishable results on the yeast diauxic shift will be an objective evidence for the success of the project.
Investigation of the Warburg effect in cancer, and of calorie restriction in ageing using yeast as a model system. We will use yeast diauxic shift to investigate the Warburg effect. A deeper understanding of the mechanisms responsible for the diauxic shift in yeast may suggest new opportunities for therapeutic intervention in cancer, and aging.